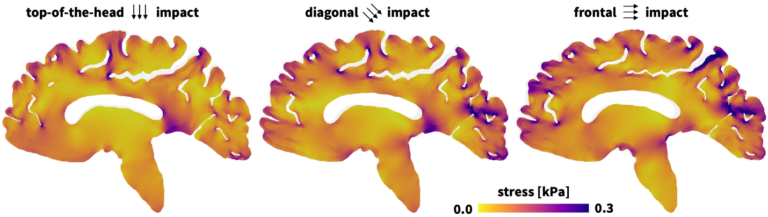
Simulation is a powerful tool that health care professionals can leverage to analyze patient data at the tissue level.
By Katie Corey
Simulation software has been a product design and engineering staple for over 40 years. It is still a practical and powerful tool that allows engineers to digitally test designs early and under different stresses and strains. It’s all about efficiency: Quickly find potential design problems to prevent costly or hazardous issues downstream and save time and money in the process.
What if researchers and health care professionals could analyze patient data at the tissue level by leveraging industrial simulation tools? A similar level of safety, reliability, and efficiency could be built into medical diagnosis, device design, and interventions using the same finite element analysis (FEA) tools that engineers have used for decades to validate and optimize motor vehicle and aircraft designs.
Expanding Biological Modeling to Study Brain Trauma
The number of studies regarding traumatic brain injury, which is particularly relevant in sports and traffic accidents, has increased in recent years. Researchers at Stanford University wanted to build a more accurate AI-based model to understand how brain deformations translate to stress and strain that potentially cause lasting brain damage. The model needed to address the issue of the brain’s heterogeneous nature and ultra-soft material makeup, which makes physical testing and modeling extremely challenging. Historically researchers relied on a variety of models for studying brain trauma, some nearly a century old. These models often applied only to specific types of stress or brain regions.
“We want researchers to be able to go from experimental data to material modeling to an FEA simulation with minimal user intervention,” explains Mathias Peirlinck, former postdoc at Stanford University and now professor at Delft University of Technology specializing in soft tissue biomechanics, computational cardiac biophysics, and machine learning.
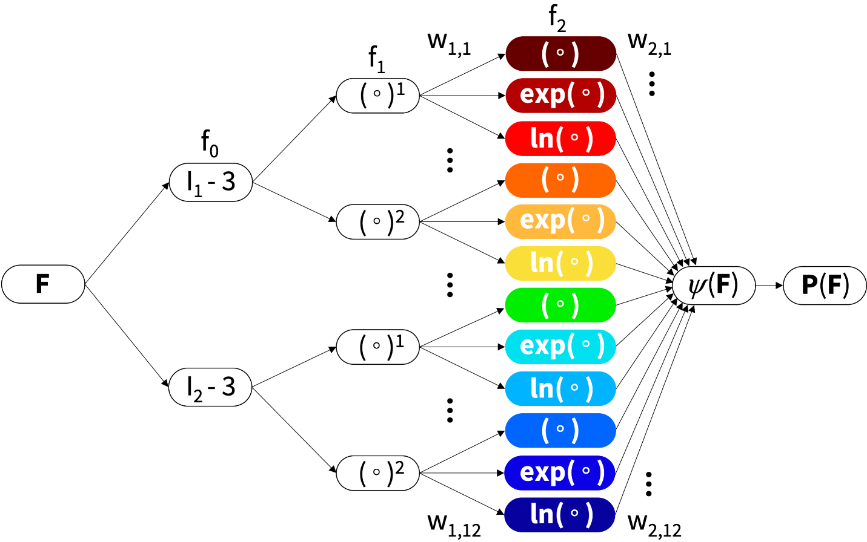
Stanford researchers developed Constitutive Artificial Neural Networks (CANNs), which leverage AI via an integration of physics and machine learning to enable automated model discovery. Developed by Kevin Linka, a former postdoc at Stanford and now a professor at RWTH Aachen, the CANN method automates the selection process from over 4,000 models. While such selection was previously based on user experience and personal preference, the network autonomously selects the best model, parameters, and experiment to characterize soft matter systems.
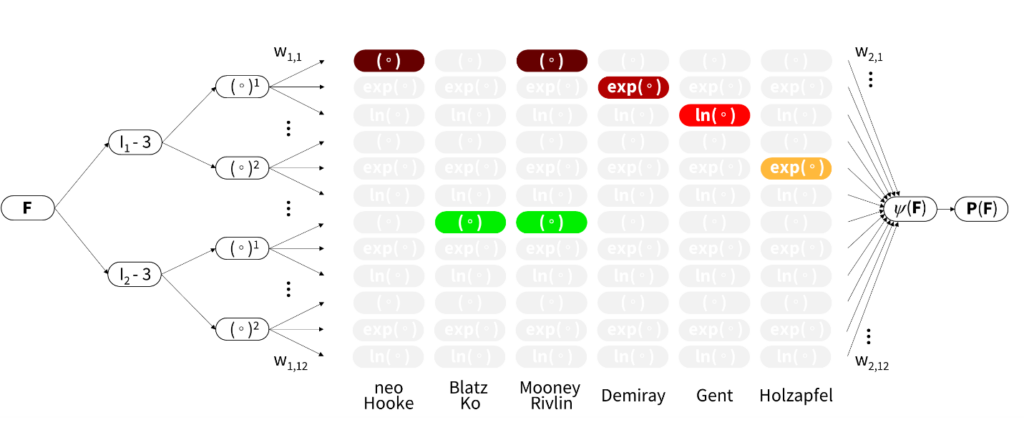
“We challenge the conventional wisdom to first select a constitutive model and then fit its parameters to experimental data,” explains Ellen Kuhl, professor of mechanical engineering at Stanford specializing in machine learning and living systems. “Instead of using classical off-the-shelf neural networks, we reverse engineer our own network from a set of functional building blocks that are, by design, a generalization of widely accepted models.”
Machine learning is a subset of AI focused on building systems that can learn from and make decisions based on data, enabling computers to improve their performance on a specific task over time without being explicitly programmed for that task.
Historically, students who attended materials classes at Stanford typically faced a steep learning curve since there are dozens of different models to choose from. The CANN method takes user selection out of the equation by enabling AI to examine the data and decide which model best reflects the clinical or experimental data. “Now students will be able to, fairly quickly, understand everything that material scientists in this field have been doing for the past fifty years,” enthuses Kuhl.
Artificial neural networks are powerful computational models that mimic how the human brain processes information, enabling machines to learn from and make decisions based on data.
After the open-source CANN system was fully operational, the Stanford University team wanted to bring discovered models of the brain into FEA software. Once imported, users could simulate myriad stress and strain scenarios in a 3D digital environment to further expand understanding of brain behavior and conduct additional studies virtually.
Long-time users of SIMULIA’s Abaqus software, Kuhl and Peirlinck approached Dassault Systèmes to discuss a possible collaboration. Kuhl already had a strong relationship with the company because she was a founding member of the Living Heart Project. Would Dassault Systèmes be willing to develop an automated way to import CANN-selected models into the Abaqus FEA software? Kuhl and Peirlinck’s wish soon materialized into a presentation to the SIMULIA team in early 2023.
Connecting to Advanced Simulation
Juan Hurtado, Structures Technology Director at Dassault Systèmes, who has been involved in developing nonlinear mechanics and materials modeling functionality in the Abaqus FEA software for over twenty years, attended that presentation. “A few weeks after that initial meeting, I basically had a vision of how this could be integrated with Abaqus, and I provided an initial prototype of a universal subroutine,” he notes. Kuhl and Peirlinck were impressed and enthusiastic about partnering with Dassault Systèmes and are further developing Hurtado’s ideas.
At the outset Hurtado noted several of the myriad challenges to be faced: The CANN was great for very simple-use cases, where you have a uniform type of deformation, but what about non-uniform deformations? How do you go from the selected model to an actual 3-D simulation? If you’re dealing with brain tissue, how do you simulate that brain under impact? How do you simulate a full heart model or an artery? What if the model you have just discovered from the CANN is not directly available in the finite element software’s material library?
“I felt that Abaqus was very well positioned to integrate the discovered model from Stanford because it has quite an open and highly customizable materials modeling interface,” explains Hurtado. Also, Abaqus already could describe anisotropic hyper elastic materials that are specifically well suited for modeling soft biological tissue, an extremely challenging type of material to model.
Anisotropic hyperelasticity describes materials that can undergo large deformations and have direction-dependent properties. The stress-strain relationship is nonlinear and varies depending on the direction of the applied force.
Hurtado and his development team integrated Stanford’s CANN solution into Abaqus by developing a “Universal Material Subroutine” that now replaces dozens of individual subroutines. The entire process—from test data to constitutive model to FEA—is streamlined to the point that non-experts can use it for their materials research. “Of course, there are certain limitations, but at the same time, we are talking about a completely new paradigm in materials modeling,” notes Hurtado.
A single material subroutine defines the behavior of a specific material under various conditions. It translates strains into stresses and calculates how the material responds to different loads, deformations, or temperatures, enabling accurate predictions in real-world scenarios.
When the constitutive model is imported and translated by the Universal Material Subroutine into something that Abaqus understands, users can take full advantage of other FEA features in Abaqus, such as personalized geometries from medical images, contact, loading, or specific boundary conditions, to analyze not only the brain, but also the heart, arteries, and other bodily tissue. In fact, anyone can now analyze entire organ systems while leveraging the same Abaqus functionality used for conducting FEA studies on industrial products.
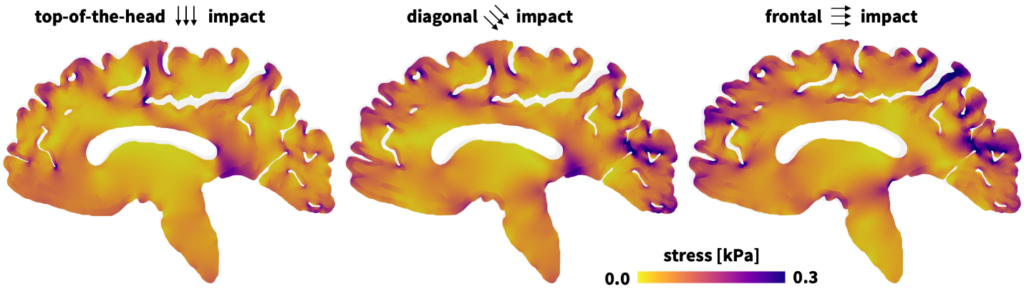
“One of the nice features of the Constitutive Artificial Neural Network is that it is constructed to produce models that obey the laws of physics. But the most powerful part is that it can generate material models that don’t currently exist—and are now included—in the Abaqus material library,” explains Hurtado. “When combined with the Universal Material Subroutine, we can validate that the response is correct for simple deformation modes and then use it for these bigger three-dimensional models of a brain.”
Moving from the Head to the Heart
The technology to study the brain that Stanford has integrated with SIMULIA’s Abaqus product is now applied to cardiovascular tissues by a team headed by Peirlinck. “I’ve worked a lot with Abaqus during my engineering studies and PhD, during my postdoc at Stanford, and now within my own research group at TU Delft,” he explains.
The most logical move, per Peirlinck, was to extend this work to the heart, which from a materials standpoint, is even more complex than the brain. Peirlinck believes that computational heart modeling is at a point where it can really benefit health care—for both patients and physicians—by improving diagnosis, treatment planning, and medical device design. “It has been an exciting opportunity to use these approaches on cardiovascular tissues as well and then immediately translate them into biomedical simulations,” says Peirlinck. As with the brain, one of the first steps in developing heart models or digital twins of organs in the human body is to choose the material model that best describes the tissue. This data varies from patient to patient, from young to old, and from healthy to diseased.
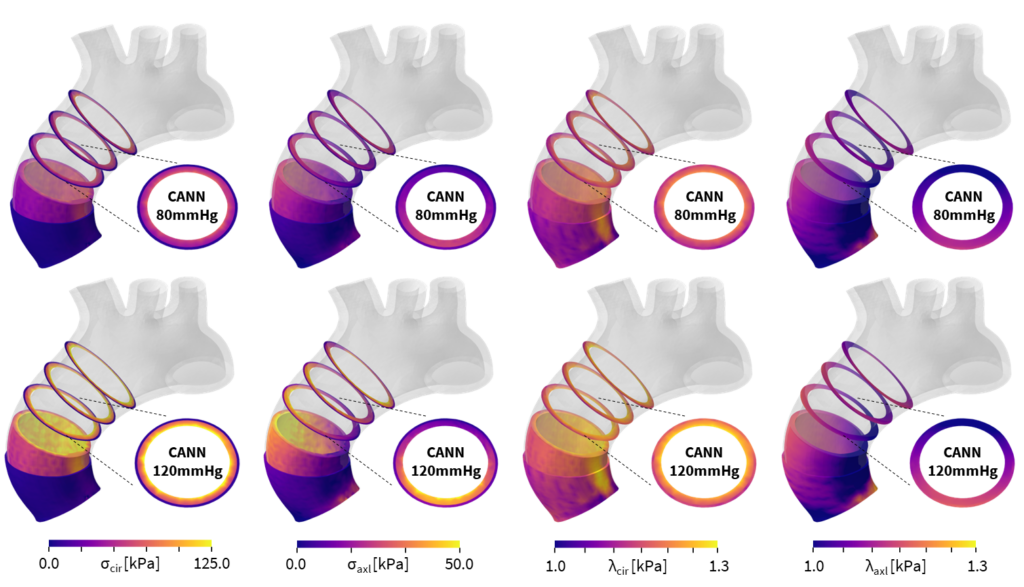
Before working with the Abaqus team, Peirlinck had to write his own code if a discovered model did not exist in the material library. After collaborating with Hurtado and the Abaqus development team, the Universal Material Subroutine guarantees that every model discovered by the CANN can be seamlessly implemented into Abaqus. “Abaqus always did a good job of bringing in the latest new constitutive models that were out there. However, now we are not limited by the material library because you can basically use any new model that you discover and then automatically integrate it into your FEA pipeline,” enthuses Peirlinck. “I think this will democratize the usage of finite element analysis for less experienced people like our medical collaborators.”
A constitutive model describes the relation between stress and strain in a material, which is essential for understanding and predicting the behavior of materials under different loading conditions—how a material deforms (strain) in response to applied forces (stress).
By leveraging this new method, users can model anything they can think of, even foams, hydrogels, or any other type of soft matter. “There are so many tissues in our body. There’s liver, there’s cartilage, there’s tendon, there’s muscle. All these have their own specific behavior,” explains Peirlinck. “The key thing that we, as engineers, need to do is get an accurate model that predicts and generalizes things we have not yet seen. We can now virtually test all these materials, tissues, and matter within the FEA analysis.” Peirlinck looks forward to how these innovations will help improve human health care by giving researchers a much clearer understanding of biological tissues.
Discovering A Better Future for Health Care
As a result of the collaboration with Dassault Systèmes, Stanford already has several success stories, including:
- Discovery of a new model for the brain that describes human gray and white matter tissue better than any previous model.
- Discovery of a first-ever material model for artificial meat by a class of students totally untrained in Abaqus.
- More than 50,000 discoverable material models that translate automatically into an Abaqus input file that forms the interface with the new Universal Material Subroutine, which will replace dozens of individual material subroutines and include thousands of new models.
“We are always speaking with clinicians about our work since we are continually translating our models into clinical situations,” says Peirlinck. “We are working with innovators around the world to build personalized models. In the future, we could envision having a quantitative physics-based model of your heart, for example, which computes, in real time, what would be the best device to treat a leaking valve. More data for better clinical decisions will translate to better patient outcomes.”
Peirlinck emphasizes that the goal isn’t to take over the clinician’s role but to provide additional data to aid the clinician with more insights to make better-informed decisions. “We have been building and improving these models for over the last decade at least, trying to get closer and closer to biological and biophysical reality. Now, with automated model discovery, more individuals can participate in this research and development. That is why this kind of automation is important,” Peirlinck adds.
Kuhl concludes, “For more than two decades, Abaqus has pushed the frontiers in material modeling both in industry and academia. We are super excited to work with Juan Hurtado and his team to integrate it with machine learning and bring material modeling to a whole new level.”